Note: The sequences on this page may not be the same length as those in the manuscript due to the end trim that was done in order to effectively BLAST these sequences. As a result, the sequences might appear to be different than those in the manuscript from the originating labs if they trimmed the raw data at a different place in the sequence. The BLAST results are the same though.
Phylogenetic tree assembled by the University of Texas for the mitochondrial DNA from Sample 25/26 that was derived from the whole genome.
Results of PP16 DNA profile for 25/26. Note the sample is male and all of the lab personnel that tested this sample were female (except for histopathology which is not genetic testing). This profile also clearly excludes the submitter's profile that is posted on another website.
This page shows some of the supporting data and results directly received from outsource laboratories used in this study. This will include raw data from the sequencers as well as some summary information prepared and sent by the outsource laboratories. For clarification purposes, Sample 25 is an extraction performed robotically by the North Louisiana Criminalistics Laboratory from 3 hairs plucked from the Northern California tissue sample labeled 26 in the study. Sample 26 was a 3 mm cube of tissue derived from the center of the same tissue as sample 25 and extracted by DNA Diagnostics. Both samples were run in tandem with identical results. You will note this in the summary chart. This data is intended to put to rest any questions about our study and the data quality. This is just a sample of what we saw consistently in all of the samples tested in the study. All testing was performed with human controls, up to 5 of them at times. The human controls all amplified and sequenced as expected on all platforms
Partial charts prepared by accredited Laboratories used in the Sasquatch Genome Project to show overall results for samples 25/26 and 31.
Below chromatogram showing 800 plus bases of sequence for the MC1R gene derived from Sample 25/26. No degradation is present with this long sequence from a single amplicon.
BLAST results for sequence for the MC1R gene derived from Sample 25/26.
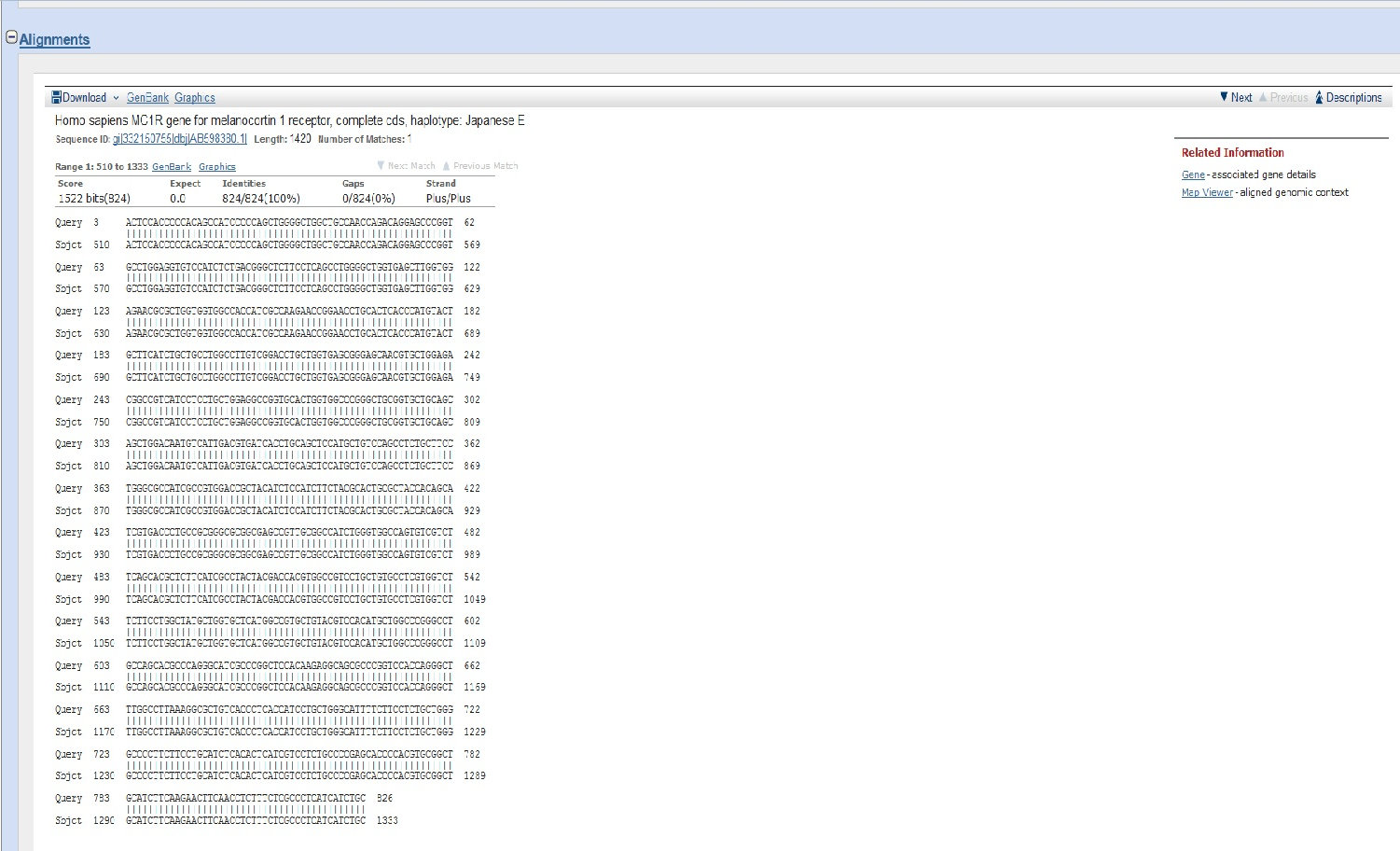
MC1R Sequence for Sample 25/26
This page shows some of the supporting data and results directly received from outsource laboratories used in this study. This will include raw data from the sequencers as well as some summary information prepared and sent by the outsource laboratories. For clarification purposes, Sample 25 is an extraction performed robotically by the North Louisiana Criminalistics Laboratory from 3 hairs plucked from the Northern California tissue sample labeled 26 in the study. Sample 26 was a 3 mm cube of tissue derived from the center of the same tissue as sample 25 and extracted by DNA Diagnostics. Both samples were run in tandem with identical results. You will note this in the summary chart. This data is intended to put to rest any questions about our study and the data quality. This is just a sample of what we saw consistently in all of the samples tested in the study. All testing was performed with human controls, up to 5 of them at times. The human controls all amplified and sequenced as expected on all platforms
TCACTCCACCCCCACAGCCATCCCCCAGCTGGGGCTGGCTGCCAACCAGACAGGAGCCCGGTGCCTGGAGGTGTCCATCTCTGACGGGCTCTTCCTCAGCCTGGGG
CTGGTGAGCTTGGTGGAGAACGCGCTGGTGGTGGCCACCATCGCCAAGAACCGGAACCTGCACTCACCCATGTACTGCTTCATCTGCTGCCTGGCCTTGTCGGACC
TGCTGGTGAGCGGGAGCAACGTGCTGGAGACGGCCGTCATCCTCCTGCTGGAGGCCGGTGCACTGGTGGCCCGGGCTGCGGTGCTGCAGCAGCTGGACAATGTCAT
TGACGTGATCACCTGCAGCTCCATGCTGTCCAGCCTCTGCTTCCTGGGCGCCATCGCCGTGGACCGCTACATCTCCATCTTCTACGCACTGCGCTACCACAGCATCGT
GACCCTGCCGCGGGCGCGGCGAGCCGTTGCGGCCATCTGGGTGGCCAGTGTCGTCTTCAGCACGCTCTTCATCGCCTACTACGACCACGTGGCCGTCCTGCTGTGCCT
CGTGGTCTTCTTCCTGGCTATGCTGGTGCTCATGGCCGTGCTGTACGTCCACATGCTGGCCCGGGCCTGCCAGCACGCCCAGGGCATCGCCCGGCTCCACAAGAGGCA
GCGCCCGGTCCACCAGGGCTTTGGCCTTAAAGGCGCTGTCACCCTCACCATCCTGCTGGGCATTTTCTTCCTCTGCTGGGGCCCCTTCTTCCTGCATCTCACACTCAT
CGTCCTCTGCCCCGAGCACCCCACGTGCGGCTGCATCTTCAAGAACTTCAACCTCTTTCTCGCCCTCATCATCTGC
Chromatogram showing sequence for the PNLIP gene derived from Sample 25/26
BLAST results for sequence for the PNLIP gene derived from Sample 25/26.
PNLIP Sequence for Sample 25-26
GTAGTAGGGTGAGGACTCTCAATGCTTCATTACCATTTGAACAGCTTCTTGTCAAGCCTGCAATCCTGAGATGTGAGGAGTGACAGAGGCTGGTAGAAAGTAGGGTTT
TGATGGGAGCTCCAATCTGCCTCTGGTGTTGCAGGGTCTCCTATGTATCAAGAACATTGAGCTGCTCCAAGTTCAGATCAGTTGTGTCTAGGGACCCAGG
Chromatogram for the M16 gene derived from Sample 25/26.
M16 Sequence for Sample 25-26
TTTTAAGTGAATAAACTTTAAACAAATACCTCCCTCGTGGCAGCAGTCTACAGTGTTTGCACGGAATGTGATCACATCCATCTCTGGGCTTAAGTCACCTGAGGTCAG
ACAGACAAGCTGATGACCACCCTCCATAGCCGCACCCCATTTTGTCCGCTGTATTATCCCCAATGAGTTTAAGCAATCGGGTATGTTGTGGGGTCACGGCTCCCTGC
TGGAAGTCCTGAGGTTCTGT
Chromatogram for the Amel X gene and associated BLAST derived from Sample 25/26.
Sequence alignments from Amel X BLAST. This one amplicon for Sample 25/26 contained loosely aligning sequences from both dog and human. Therefore, it is unknown sequence. When looking at the BLAST chart, you will notice that there is a somewhat aligned sequence to the left. That sequence is very loosely aligned with dog but there are many bases (SNPs) within that sequence that do not align with dog as seen in the sequence alignments generated by the BLAST (alignments below). Then you will see a gap in the alignment chart which shows sequence that doesn't align with anything. Then to the right you will see a sequence that aligns loosely with human. However, once again there are many bases (SNPs) within that sequence that are not consistent with human. So the sequence is not human. It is the only way that GenBank can deal with unknown sequences.
Amel X Sequence for 25-26
GCAGGGGGCCCGATGTGAGACTCGATCCTGGGACTCCAGGATCACACTCTGGATCAAAGGCAGGTGCTAAACTGATGAACCACCCAGGGATCCCTAGAGAGCACTTTC
TGAATTGTAAGGATTATATTTGTTTATTTCCTATCCCCTCTCTTCACCCAGAAAAGGATTATAGCAGTCTTTGGACAGC
Chromatogram for the Amel Y, Exon 2 derived from Sample 25/26.
Chromatogram for the Amel X gene and associated BLAST derived from Sample 25/26.
Sequence alignments from AmelY Exon 2 BLAST below. This one amplicon for 25/26 contained loosely aligning sequences from a newly uploaded sequence for a European Polecat. The previous closest aligning sequence was 70% Rhesus Monkey, now it is 86% European Polecat due to the newly uploaded Polecat sequence. However, it is unknown sequence. There are many single base mutations within that sequence that do not align as seen in the sequence alignment generated by the BLAST below. Also, the sequences are not on the chromosome so it is purely luck that there was some percentage of alignment. GenBank BLASTs select the closest aligning sequences when dealing with unknown sequences. It does not mean there is European Polecat in this sequence or Rhesus monkey, only that those are the closest sequences to the unknown sequence.
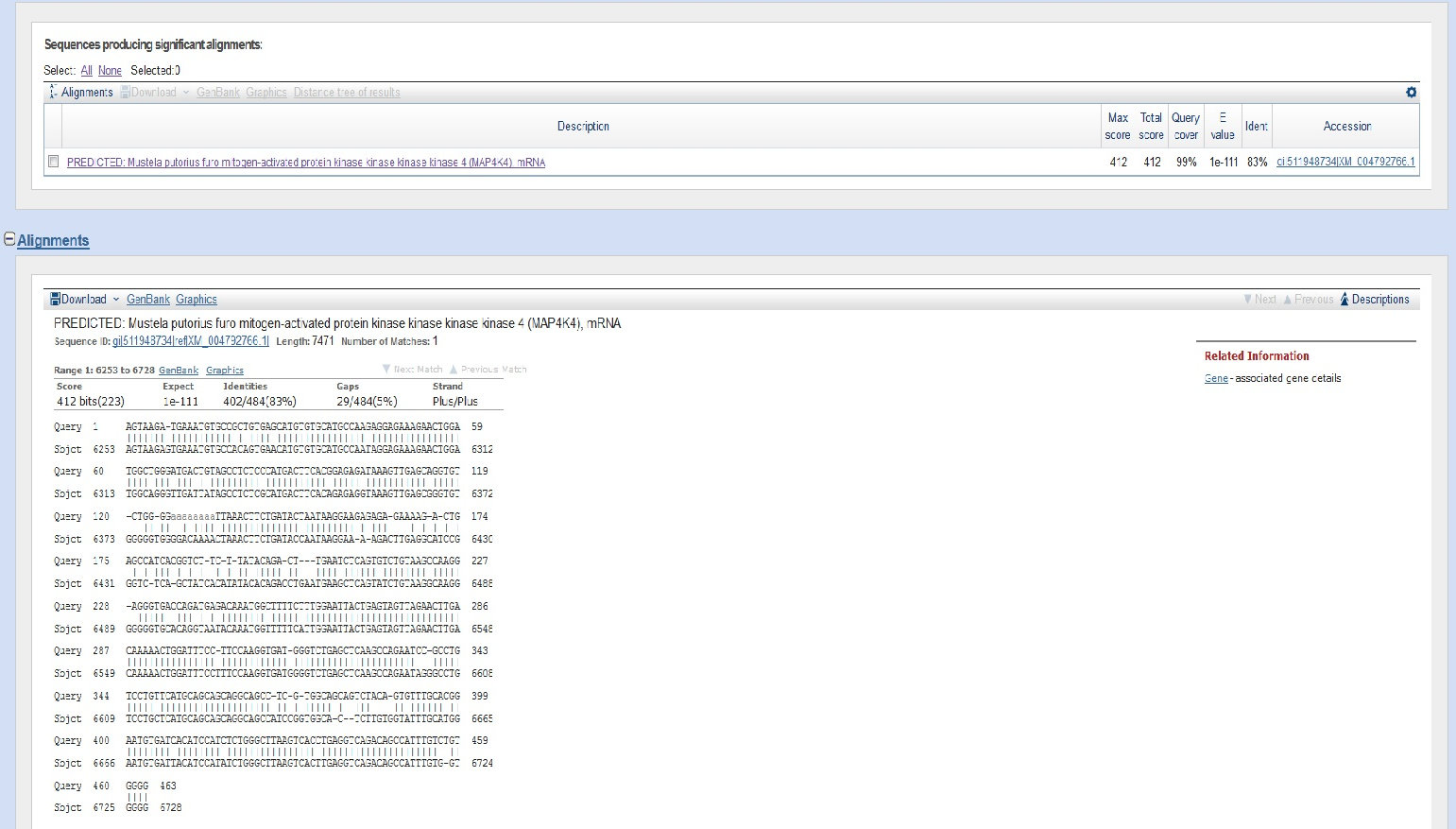
AGTAAGATGAAATGTGCCGCTGTGAGCATGTGTGCATGCCAAGAGGAGAAAGAACTGGATGGCTGGGATGACTGTAGCCTCTCCCATGACTTCACGGAGAGATAAAGTTGA
GCAGGTGTCTGGGGAAAAAAAATTAAACTTCTGATACTAATAAGGAAGAGAGAGAAAAGACTGAGCCATCACGGTCTTCTTATACAGACTTGAATCTCAGTGTCTGTAAGCC
AAGGAGGGTGACCAGATGAGACAAATGGCTTTTCTTTGGAATTACTGAGTAGTTAGAACTTGACAAAAACTGGATTTCCTTCCAAGGTGATGGGTCTGAGCTCAAGCCAGAA
TCCGCCTGTCCTGTTCATGCAGCAGCAGGCACCTCGTGGCAGCAGTCTACAGTGTTTGCACGGAATGTGATCACATCCATCTCTGGGCTTAAGTCACCTGAGGTCAGACAGG
CCATTTGTCTGTGGGGTA
Amel Y, Exon 2, Sequence for 25-26
Chromatogram and BLAST for HAR1 derived from Sample 25/26. Completely unknown sequence.
HAR 1 Sequence for Sample 25-26
ATCTGAAATGTGAGACCCAGGGAAAGAGAATGATATTCTACCAAATAACAGGTCTGTAATCTTTAAAAACTGGTCGTAAAGGTCAGGGAAAGGCGGTGGAACGGTTTCA
GATCACGGAGACTAACGAGGCTTGAGTGGTTGAACCGGACCCTCTCCGTGGGGAGGACGGGCCGCTCCCGAGTGGGTGTGAGGACTGGCCCGCCGCACAGTAACA
AGCTACCTGCCCGCTCCTGGTGGCCGCTTAGGTGGCATA
DOWNLOADS:
Supplemental Raw Data

